Principles of gene regulation:
Introduction to Gene Regulation
Gene regulation is a critical process by which cells control the expression of genes to ensure proper function, adaptation, and development. In every organism, genes encode information that guides cellular activities, but not all genes are expressed at all times or in all cells. Gene regulation ensures that genes are turned on or off in response to specific signals, environmental changes, or developmental cues, which is essential for maintaining cellular homeostasis, differentiation, and response to external factors.
In eukaryotes, gene regulation allows for complex processes such as tissue-specific gene expression, the coordination of metabolic pathways, and the response to stress or signaling molecules. In prokaryotes, gene regulation is vital for adapting to changes in the environment, such as the availability of nutrients. This regulation occurs at various levels, from the DNA to the protein, and involves multiple mechanisms, including modifications to the DNA itself, RNA processing, and changes in protein activity.
The study of gene regulation is foundational to understanding how cells function, how organisms develop, and how diseases such as cancer and genetic disorders arise due to misregulation of genes. This broad field also has significant applications in biotechnology, where manipulating gene expression can be used for medical therapies, agricultural improvements, and industrial applications.
In this context, gene regulation can be categorized into several key areas, each playing a vital role in determining when, where, and how genes are expressed. These mechanisms ensure that cellular processes are finely tuned and that gene expression is responsive to both intrinsic and extrinsic factors. The following sections will explore these regulatory mechanisms in detail, including the various levels at which gene expression is controlled and how these processes contribute to cellular function and organismal development.
Reference: libretexts Biology
1. Types of Gene Regulation
Gene regulation occurs at several levels, from the transcription of DNA into RNA to the eventual synthesis and function of proteins. The regulation can be broadly categorized into the following types:
Transcriptional regulation: This is the most fundamental level of gene regulation and involves controlling the process by which mRNA is synthesized from DNA.
Post-transcriptional regulation: After transcription, mRNA can undergo processing events that control its stability, splicing, export from the nucleus, and eventual degradation.
Translational regulation: At this level, the translation of mRNA into protein is controlled. This can involve regulating the initiation of translation, as well as the stability and activity of the mRNA.
Post-translational regulation: After a protein is synthesized, it can undergo modifications such as phosphorylation, acetylation, and ubiquitination that affect its function, localization, or stability.
2. Gene Regulation at the Transcriptional Level
Transcriptional regulation controls the synthesis of RNA from a gene's DNA template. It is crucial in controlling when and how much mRNA is produced. Key elements include:
Promoter Region
The promoter is a DNA sequence located near the gene that serves as the binding site for RNA polymerase, the enzyme that synthesizes RNA. Transcription factors help recruit RNA polymerase to the promoter, and the strength or accessibility of the promoter can determine the level of gene expression.
Transcription Factors
Transcription factors are proteins that bind to specific DNA sequences to regulate the transcription of a gene. There are two types:
- Activators: These enhance transcription by binding to the promoter or enhancer regions, recruiting RNA polymerase or coactivators.
- Repressors: These inhibit transcription by binding to operator regions or silencer sequences, preventing RNA polymerase from initiating transcription.
Enhancers and Silencers
Enhancers are DNA sequences that can increase the transcription of a gene. They are often located far away from the gene they regulate, sometimes even on different chromosomes. Enhancers work by being bound by activators or coactivators, which bend the DNA to bring the enhancer closer to the gene promoter.
Silencers, in contrast, are DNA sequences that suppress gene expression by binding repressor proteins that block transcription.
Operons (in prokaryotes)
An operon is a cluster of genes that are transcribed together from a single promoter and operator. In bacteria, operons are used to regulate the expression of related genes in response to environmental changes. A classic example is the lac operon in E. coli, which regulates genes involved in lactose metabolism. The lac operon is repressed when lactose is not available, and activated when lactose is present, allowing efficient use of resources.
3. Epigenetic Regulation
Epigenetics involves modifications to the DNA or histones that affect gene expression without altering the DNA sequence itself. These changes can be inherited and influence gene expression over generations.
DNA Methylation
DNA methylation involves the addition of a methyl group (-CH3) to the DNA, usually at cytosine residues in CpG dinucleotides. Methylation of DNA typically represses gene expression by preventing the binding of transcription factors and making the DNA less accessible. Aberrant DNA methylation is often observed in diseases like cancer.
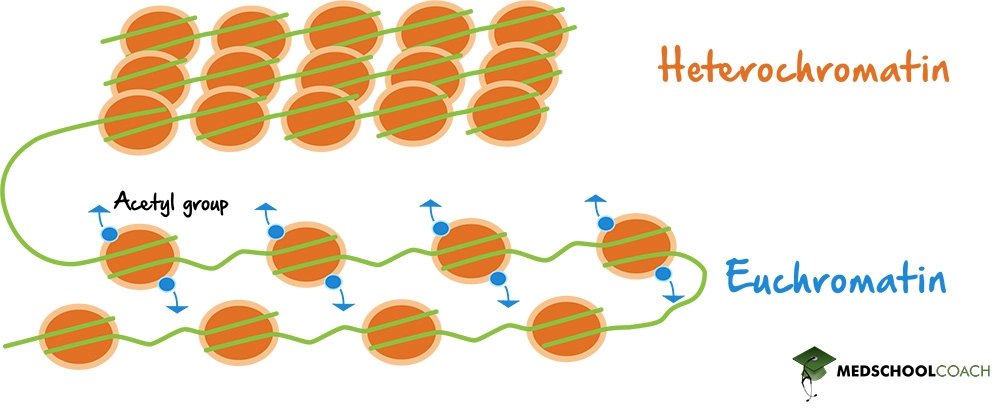
Histone Modification
Histones are proteins around which DNA is wound to form chromatin. Chemical modifications to histones, such as acetylation, methylation, phosphorylation, and ubiquitination, can affect chromatin structure and gene accessibility. Histone acetylation generally correlates with gene activation, while histone methylation can be associated with both activation or repression, depending on the context.
Chromatin Remodeling
Chromatin remodeling complexes can reposition or restructure nucleosomes (the basic unit of chromatin) to make DNA more or less accessible to the transcriptional machinery. This is an important regulatory mechanism for allowing or preventing transcription initiation.
4. Post-Transcriptional Regulation
After transcription, the resulting mRNA transcript can be modified, stabilized, or degraded. Several key mechanisms are involved:
RNA Splicing
Eukaryotic genes are often interrupted by non-coding regions called introns, which must be removed during the splicing process. The remaining coding regions, called exons, are spliced together to form mature mRNA. Alternative splicing allows a single gene to produce multiple protein isoforms by selecting different combinations of exons.
RNA Stability and Degradation
The half-life of mRNA in the cell determines how long it can be translated into protein. Regulatory proteins and non-coding RNAs (e.g., microRNAs) can bind to mRNA molecules, marking them for degradation. The process of mRNA decay is crucial for controlling gene expression, especially under stress conditions.
RNA Interference (RNAi)
Small RNA molecules, such as microRNAs (miRNAs) and small interfering RNAs (siRNAs), can bind to mRNA molecules and either degrade them or inhibit their translation. RNAi is a potent mechanism for regulating gene expression in response to developmental cues, stress, and viral infections.
RNA Editing
RNA editing refers to modifications made to the RNA molecule itself, often involving the conversion of one nucleotide to another. For example, adenosine can be converted to inosine, which is read as guanine by the ribosome, potentially altering the protein produced. RNA editing contributes to the diversity of proteins in a cell.
5. Translational Regulation
Translational regulation controls the efficiency and rate at which mRNA is translated into protein.
Regulation of Translation Initiation
The initiation of translation can be regulated by the availability of translation initiation factors and ribosomal components. In many cases, the presence of specific regulatory proteins or RNAs can inhibit the binding of ribosomes to the mRNA, thereby preventing translation.
mRNA Structure
The 5' untranslated region (UTR) of mRNA often contains elements that regulate translation. For example, some sequences in the 5' UTR may bind to regulatory proteins or small RNAs that block or promote translation initiation.
Regulation by RNA Binding Proteins
RNA-binding proteins can bind to specific sites on mRNA and either enhance or inhibit translation. These proteins are often regulated by signaling pathways that respond to environmental conditions, stress, or developmental signals.
6. Post-Translational Regulation
Once proteins are synthesized, their activity and stability can be further regulated by several mechanisms:
Protein Folding and Chaperones
After translation, proteins must fold into their functional three-dimensional structure. Molecular chaperones assist in this process, ensuring proteins fold correctly. Misfolded proteins are often degraded to maintain cellular integrity.
Phosphorylation
Phosphorylation is the addition of phosphate groups to proteins, typically on serine, threonine, or tyrosine residues. This modification can activate or inactivate enzymes, change protein localization, or alter protein-protein interactions.
Ubiquitination
Ubiquitination involves the attachment of small ubiquitin molecules to proteins, marking them for degradation by the proteasome. This process helps regulate protein levels and ensures the removal of damaged or unneeded proteins.
Acetylation and Methylation
Like histones, non-histone proteins can undergo acetylation and methylation, which can impact their function. For example, acetylation of transcription factors can activate their DNA-binding activity, whereas methylation can repress their function.
7. Feedback Loops in Gene Regulation
Feedback loops help to maintain homeostasis and regulate gene expression in response to changing internal and external conditions.
Positive Feedback
In a positive feedback loop, the product of a gene expression amplifies its own production. This can create a self-perpetuating cycle of gene expression. For example, the transcription factor Myc can promote its own expression, leading to the enhanced expression of genes involved in cell cycle progression.
Negative Feedback
In negative feedback, the product of a gene expression inhibits its own production. For example, a transcription factor that is synthesized in response to a signal may later inhibit its own expression, preventing overactivation of a pathway.
8. Gene Regulation in Eukaryotes vs Prokaryotes
Prokaryotes (e.g., E. coli)
In prokaryotes, gene regulation is simpler. Genes are often organized into operons, which are regulated together by a single promoter and operator region. Prokaryotes use repressors and activators to regulate gene expression in response to environmental signals, such as nutrient availability.
Eukaryotes
Eukaryotic gene regulation is more complex due to the compartmentalization of transcription (in the nucleus) and translation (in the cytoplasm). Eukaryotes also have more sophisticated mechanisms such as chromatin remodeling, alternative splicing, and post-transcriptional modifications. Gene expression in eukaryotes is highly regulated in response to signals during development, differentiation, and environmental changes.
Please, give your suggestions in comment section.
Follow our blogs for more updates and protocols: https://iammolecularbiologist.blogspot.com/
Follow us on WhatsApp: https://chat.whatsapp.com/I30OdffWlJhGABXsTmKo95