DNA Packaging:
DNA packaging is a highly regulated process that enables the compaction of genetic material to fit within the confines of the cell nucleus while still allowing access for essential biological processes such as transcription, replication, and repair. This organization involves a hierarchical structure, from the basic nucleosome unit to higher-order chromatin structures.
1. Nucleosome Formation
Histones and DNA Interaction:
- Histone Proteins: DNA is first wrapped around histone proteins to form the basic unit of chromatin, the nucleosome. Histones are small, highly basic proteins that neutralize the negative charge of DNA through electrostatic interactions.
- Histone Octamer: A nucleosome core particle consists of an octamer of histone proteins (two each of H2A, H2B, H3, and H4). The DNA, approximately 146 base pairs in length, wraps around this octamer about 1.7 times, forming a histone core.
- Histone H1: The linker histone H1 binds to the entry and exit sites of DNA on the nucleosome, stabilizing the nucleosome structure and promoting higher-order chromatin formation.
Structural Characteristics:
- Bead-on-a-String Model: The initial structure of chromatin is often described as a "bead-on-a-string" model, where nucleosomes (beads) are connected by linker DNA (string).
2. Formation of Higher-Order Chromatin Structures
30 nm Fiber:
- Chromatin Fiber Formation: The "bead-on-a-string" nucleosomal array is further compacted into a 30 nm fiber. This fiber can be organized into two main models: the solenoid model (a helical structure) and the zigzag model (a less regular, more extended structure).
- Compaction: The 30 nm fiber is believed to be stabilized by interactions between histone tails and linker DNA.
Loop Domains and Chromatin Organization:
- Loop Domains: The 30 nm fiber is organized into looped domains anchored to a nuclear matrix or scaffold. This organization involves a network of structural maintenance of chromosomes (SMC) proteins, such as cohesin and condensin.
- Chromosome Territories: In the nucleus, individual chromosomes occupy distinct territories. This organization influences gene expression and chromatin accessibility by reducing interactions between chromosomes.
3. Histone Modifications and Chromatin Remodeling
Post-Translational Modifications:
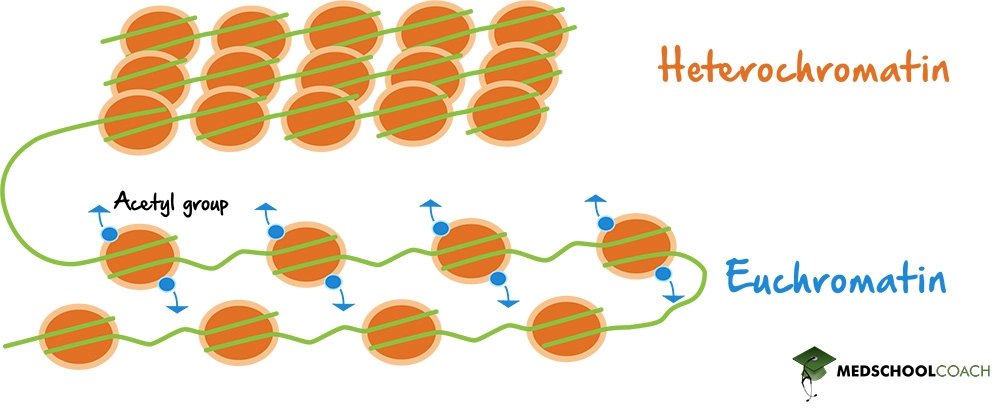
- Acetylation: Histone acetylation, mediated by histone acetyltransferases (HATs), adds acetyl groups to lysine residues, reducing their positive charge and decreasing histone-DNA interactions. This modification is generally associated with gene activation.
- Methylation: Histone methylation can either activate or repress gene expression depending on the context. Methylation of H3K4 is often associated with active transcription, while methylation of H3K9 and H3K27 is linked to gene repression.
- Phosphorylation: Phosphorylation of histone H3, particularly at serine 10, is associated with chromosome condensation during mitosis and is also involved in DNA damage response.
Chromatin Remodeling Complexes:
- ATP-Dependent Remodeling: Chromatin remodeling complexes, such as SWI/SNF and ISWI, use ATP hydrolysis to reposition, eject, or restructure nucleosomes, thereby influencing chromatin accessibility and gene expression.
4. Chromatin Types: Euchromatin vs. Heterochromatin
Euchromatin:
- Characteristics: Euchromatin is less densely packed and is associated with active gene transcription. It has a more open chromatin structure that allows access to the transcriptional machinery.
- Role in Transcription: Euchromatin regions are generally rich in gene-dense areas and are frequently associated with active genes and regulatory elements.
Heterochromatin:
- Characteristics: Heterochromatin is more compact and transcriptionally inactive. It can be classified into:
- Constitutive Heterochromatin: Permanently compacted regions, such as centromeres and telomeres, involved in maintaining chromosome stability.
- Facultative Heterochromatin: Can switch between an active and inactive state depending on cellular conditions. Examples include the X chromosome inactivation in females.
5. DNA Packaging in Prokaryotes
Nucleoid Region:
- Organization: Prokaryotic DNA is organized in the nucleoid region, which lacks a defined membrane-bound nucleus. DNA is associated with various non-histone DNA-binding proteins.
- Supercoiling: Prokaryotic DNA is typically negatively supercoiled, which introduces tension into the DNA, aiding in its compaction and playing a role in replication and transcription.
DNA Binding Proteins:
- Structural Maintenance: Proteins such as HU (Histone like DNA binding proteins) and IHF (Integration host factors) help in organizing DNA into loops and domains within the nucleoid, affecting gene expression and DNA replication.

6. Chromosome Condensation During Mitosis
Chromosome Structure:
- Condensation: During cell division, chromatin undergoes further condensation to form distinct, visible chromosomes. This process is facilitated by condensin complexes, which help in the compaction of chromatin and ensure proper segregation of chromosomes.
- Mitotic Chromosomes: The fully condensed mitotic chromosomes are highly compacted to ensure that DNA is accurately divided between daughter cells during mitosis.
Functional Implications
Gene Regulation:
- Transcriptional Control: DNA packaging affects the accessibility of genes for transcription. Modifications and higher-order structures can either promote or inhibit gene expression based on the cellular context.
Genomic Stability:
- Chromosome Integrity: Proper DNA packaging is crucial for maintaining genomic stability. Errors in chromatin organization can lead to genomic instability and contribute to various diseases, including cancer.
Please, give your suggestions in comment section.
Follow our blogs for more updates and protocols: https://iammolecularbiologist.blogspot.com/