Which of the following is NOT a type of RNA?
- A) mRNA
- B) tRNA
- C) rRNA
- D) dRNA
- Answer: D
What is the primary function of mRNA?
- A) Transporting amino acids
- B) Carrying genetic information from DNA to the ribosome
- C) Catalyzing chemical reactions
- D) Forming the structural components of ribosomes
- Answer: B
tRNA molecules are responsible for:
- A) Synthesizing RNA
- B) Bringing amino acids to the ribosome
- C) Storing genetic information
- D) Catalyzing peptide bond formation
- Answer: B
Which base is not found in RNA?
- A) Adenine
- B) Thymine
- C) Cytosine
- D) Uracil
- Answer: B
What sugar is found in RNA?
- A) Ribose
- B) Deoxyribose
- C) Glucose
- D) Fructose
- Answer: A
In RNA, adenine pairs with:
- A) Thymine
- B) Cytosine
- C) Guanine
- D) Uracil
- Answer: D
The backbone of an RNA molecule consists of:
- A) Nucleotide bases only
- B) Ribose and phosphate groups
- C) Amino acids
- D) Deoxyribose and phosphate groups
- Answer: B
Which of the following statements about RNA is TRUE?
- A) RNA is double-stranded
- B) RNA contains the sugar deoxyribose
- C) RNA contains uracil instead of thymine
- D) RNA is more stable than DNA
- Answer: C
The enzyme that synthesizes RNA from a DNA template is:
- A) DNA polymerase
- B) RNA polymerase
- C) Reverse transcriptase
- D) Ligase
- Answer: B
Ribosomal RNA (rRNA) is a component of:
- A) Ribosomes
- B) Mitochondria
- C) Lysosomes
- D) Golgi apparatus
- Answer: A
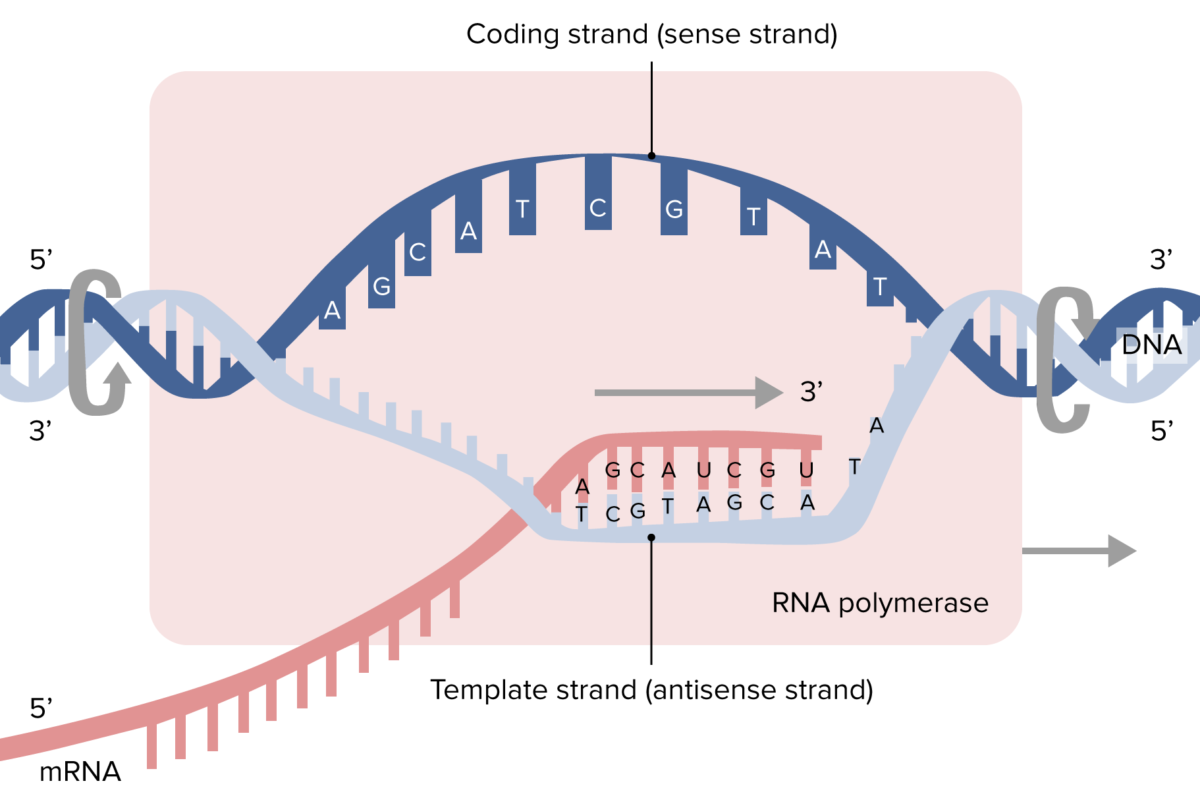
Transcription
Transcription is the process of:
- A) Replicating DNA
- B) Synthesizing RNA from a DNA template
- C) Translating RNA into protein
- D) Splicing RNA
- Answer: B
The promoter region is:
- A) A sequence where transcription begins
- B) A sequence where translation begins
- C) A sequence that codes for proteins
- D) A sequence that signals the end of transcription
- Answer: A
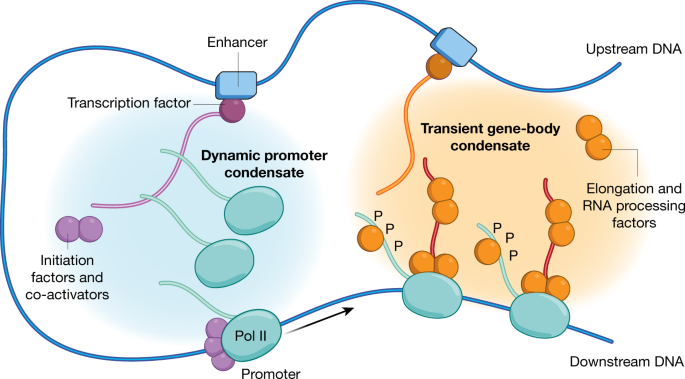
Which of the following is required for transcription initiation in eukaryotes?
- A) RNA polymerase
- B) Transcription factors
- C) Promoter region
- D) All of the above
- Answer: D
The TATA box is:
- A) A type of ribosome
- B) A sequence in the promoter region
- C) An enzyme
- D) A sequence in the terminator region
- Answer: B
In eukaryotes, transcription occurs in the:
- A) Cytoplasm
- B) Nucleus
- C) Ribosomes
- D) Mitochondria
- Answer: B
The strand of DNA that is used as a template for RNA synthesis is called the:
- A) Coding strand
- B) Template strand
- C) Leading strand
- D) Lagging strand
- Answer: B
In prokaryotes, the sigma factor is important for:
- A) RNA splicing
- B) DNA replication
- C) Transcription initiation
- D) Protein synthesis
- Answer: C
Termination of transcription in prokaryotes often involves:
- A) A stop codon
- B) A rho factor
- C) A start codon
- D) RNA polymerase binding
- Answer: B
Splicing of pre-mRNA involves:
- A) Removal of introns
- B) Addition of a 5' cap
- C) Addition of a poly-A tail
- D) All of the above
- Answer: D
Which of the following modifications occur at the 3' end of eukaryotic mRNA?
- A) 5' capping
- B) Polyadenylation
- C) Splicing
- D) Methylation
- Answer: B
RNA Function
The function of rRNA is to:
- A) Transfer amino acids
- B) Encode genetic information
- C) Catalyze peptide bond formation
- D) Regulate gene expression
- Answer: C
Small nuclear RNA (snRNA) is involved in:
- A) Protein synthesis
- B) DNA replication
- C) RNA splicing
- D) RNA degradation
- Answer: C
MicroRNAs (miRNAs) function primarily by:
- A) Catalyzing metabolic reactions
- B) Regulating gene expression
- C) Synthesizing DNA
- D) Repairing DNA
- Answer: B
Which type of RNA carries amino acids to the ribosome during translation?
- A) mRNA
- B) tRNA
- C) rRNA
- D) miRNA
- Answer: B
The anticodon is a feature of:
- A) mRNA
- B) tRNA
- C) rRNA
- D) snRNA
- Answer: B
Ribozymes are:
- A) Proteins that catalyze RNA splicing
- B) RNA molecules that act as enzymes
- C) DNA molecules that act as enzymes
- D) Proteins that synthesize RNA
- Answer: B
The wobble hypothesis is associated with:
- A) mRNA stability
- B) tRNA anticodon flexibility
- C) rRNA structure
- D) DNA replication
- Answer: B
Which of the following is a function of RNA interference (RNAi)?
- A) DNA replication
- B) Transcription initiation
- C) Gene silencing
- D) Protein synthesis
- Answer: C
Long non-coding RNAs (lncRNAs) are involved in:
- A) Coding for proteins
- B) Regulating gene expression
- C) DNA replication
- D) Translation
- Answer: B
Which RNA type is most directly involved in translation?
- A) mRNA
- B) tRNA
- C) rRNA
- D) All of the above
- Answer: D
Advanced RNA Topics
RNA editing can result in:
- A) Changes in the RNA sequence after transcription
- B) Changes in the DNA sequence
- C) Protein degradation
- D) RNA splicing
- Answer: A
The poly-A tail of mRNA:
- A) Is added during transcription
- B) Is important for mRNA stability
- C) Helps initiate translation
- D) Is found in tRNA
- Answer: B
Which of the following processes involves RNA-dependent RNA polymerase?
- A) Transcription
- B) RNA interference
- C) Reverse transcription
- D) DNA replication
- Answer: B
Which molecule is required for the initiation of transcription in prokaryotes?
- A) RNA polymerase II
- B) DNA polymerase
- C) Sigma factor
- D) Helicase
- Answer: C
Which of the following RNA types has the longest average lifespan in eukaryotic cells?
- A) mRNA
- B) tRNA
- C) rRNA
- D) miRNA
- Answer: C
RNA secondary structure is primarily determined by:
- A) Base sequence
- B) Protein interactions
- C) Temperature
- D) Intracellular location
- Answer: A
RNA molecules can form complex secondary structures such as:
- A) Alpha helices
- B) Beta sheets
- C) Hairpins and loops
- D) Z-DNA
- Answer: C
The Shine-Dalgarno sequence is found in:
- A) Eukaryotic mRNA
- B) Prokaryotic mRNA
- C) tRNA
- D) rRNA
- Answer: B
Which enzyme is responsible for adding the poly-A tail to mRNA?
- A) RNA polymerase
- B) Poly-A polymerase
- C) DNA polymerase
- D) Helicase
- Answer: B
Which of the following is NOT a component of the eukaryotic transcription initiation complex?
- A) RNA polymerase II
- B) TATA-binding protein (TBP)
- C) Sigma factor
- D) Transcription factors
- Answer: C
RNA Processing and Post-Transcriptional Modifications
The 5' cap added to eukaryotic mRNA is important for:
- A) mRNA stability
- B) Initiation of translation
- C) Export from the nucleus
- D) All of the above
- Answer: D
Alternative splicing allows for:
- A) The production of multiple proteins from a single gene
- B) Increased mRNA stability
- C) Enhanced DNA replication
- D) Gene silencing
- Answer: A
Which process removes introns from pre-mRNA?
- A) Transcription
- B) Splicing
- C) Translation
- D) Replication
- Answer: B
The branch point sequence is important for:
- A) Transcription termination
- B) Splicing of introns
- C) Translation initiation
- D) DNA replication
- Answer: B
What is the role of small nuclear ribonucleoproteins (snRNPs) in RNA processing?
- A) They degrade mRNA
- B) They assist in splicing of pre-mRNA
- C) They synthesize RNA
- D) They add the 5' cap to mRNA
- Answer: B
RNA editing can change a codon for one amino acid to a codon for another by:
- A) Substituting one nucleotide for another
- B) Deleting nucleotides
- C) Adding nucleotides
- D) Splicing out exons
- Answer: A
During RNA interference, which molecules guide the degradation of target mRNA?
- A) Ribozymes
- B) siRNAs
- C) tRNAs
- D) rRNAs
- Answer: B
RNA polymerase II is primarily responsible for transcribing:
- A) rRNA genes
- B) tRNA genes
- C) mRNA genes
- D) miRNA genes
- Answer: C
Which RNA modification is unique to eukaryotes and not found in prokaryotes?
- A) Splicing of introns
- B) Addition of a 5' cap
- C) Polyadenylation of mRNA
- D) All of the above
- Answer: D
Which structure is essential for the translation of mRNA in prokaryotes?
- A) 5' cap
- B) Poly-A tail
- C) Shine-Dalgarno sequence
- D) Spliceosome
- Answer: C
Translation and Ribosomes
Translation is the process of:
- A) Synthesizing RNA from a DNA template
- B) Synthesizing DNA from an RNA template
- C) Synthesizing proteins from an mRNA template
- D) Synthesizing mRNA from a protein template
- Answer: C
The start codon for translation is:
- A) UAA
- B) AUG
- C) UGA
- D) UAG
- Answer: B
Ribosomes are composed of:
- A) DNA and proteins
- B) RNA and DNA
- C) RNA and proteins
- D) Proteins only
- Answer: C
The large subunit of the ribosome is responsible for:
- A) mRNA binding
- B) tRNA binding
- C) Catalyzing peptide bond formation
- D) Transcription initiation
- Answer: C
Which site on the ribosome does the initiator tRNA bind to?
- A) A site
- B) P site
- C) E site
- D) Z site
- Answer: B
The function of the A site on the ribosome is to:
- A) Bind the tRNA carrying the growing polypeptide chain
- B) Bind the tRNA carrying the next amino acid to be added
- C) Release the uncharged tRNA
- D) Bind the mRNA
- Answer: B
Which molecule is responsible for bringing amino acids to the ribosome?
- A) mRNA
- B) tRNA
- C) rRNA
- D) DNA
- Answer: B
Peptidyl transferase activity is a function of:
- A) tRNA
- B) mRNA
- C) rRNA
- D) DNA
- Answer: C
The termination of translation occurs when:
- A) A stop codon is reached
- B) The ribosome reaches the end of the mRNA
- C) The ribosome binds to the Shine-Dalgarno sequence
- D) A start codon is reached
- Answer: A
Polysomes are:
- A) Single ribosomes bound to multiple mRNA molecules
- B) Multiple ribosomes bound to a single mRNA molecule
- C) Single mRNA molecules bound to multiple ribosomes
- D) Multiple mRNA molecules bound to a single ribosome
- Answer: B
RNA Regulation and Degradation
Gene expression can be regulated at the level of:
- A) Transcription
- B) RNA processing
- C) Translation
- D) All of the above
- Answer: D
Which type of RNA is involved in gene silencing and regulation?
- A) mRNA
- B) tRNA
- C) miRNA
- D) rRNA
- Answer: C
RNA stability is often controlled by:
- A) 5' cap
- B) Poly-A tail
- C) RNA-binding proteins
- D) All of the above
- Answer: D
The degradation of mRNA involves:
- A) Removal of the 5' cap
- B) Removal of the poly-A tail
- C) Endonucleolytic cleavage
- D) All of the above
- Answer: D
Which enzyme is involved in the degradation of mRNA?
- A) RNA polymerase
- B) Ribonuclease
- C) DNA polymerase
- D) Helicase
- Answer: B
Which molecule plays a key role in RNA interference (RNAi)?
- A) mRNA
- B) siRNA
- C) rRNA
- D) tRNA
- Answer: B
The function of Dicer in RNA interference is to:
- A) Degrade target mRNA
- B) Cleave double-stranded RNA into siRNAs
- C) Synthesize RNA
- D) Export mRNA from the nucleus
- Answer: B
Argonaute proteins are essential components of the:
- A) Spliceosome
- B) Ribosome
- C) RNA-induced silencing complex (RISC)
- D) DNA replication machinery
- Answer: C
Which process converts pre-mRNA into mature mRNA?
- A) Transcription
- B) Splicing
- C) Translation
- D) Replication
- Answer: B
RNA editing can involve:
- A) Deletion of nucleotides
- B) Insertion of nucleotides
- C) Substitution of nucleotides
- D) All of the above
- Answer: D
RNA Technologies and Applications
Reverse transcription is the process of:
- A) Synthesizing RNA from a DNA template
- B) Synthesizing DNA from an RNA template
- C) Synthesizing proteins from an mRNA template
- D) Synthesizing RNA from an RNA template
- Answer: B
Which enzyme synthesizes DNA from an RNA template?
- A) RNA polymerase
- B) DNA polymerase
- C) Reverse transcriptase
- D) Helicase
- Answer: C
cDNA is:
- A) Complementary DNA synthesized from an mRNA template
- B) Circular DNA found in bacteria
- C) DNA that encodes for ribosomal RNA
- D) DNA that is transcribed into tRNA
- Answer: A
Which technique can be used to measure RNA levels in a sample?
- A) PCR
- B) RT-PCR
- C) DNA sequencing
- D) Western blotting
- Answer: B
Northern blotting is used to:
- A) Detect DNA
- B) Detect RNA
- C) Detect proteins
- D) Detect lipids
- Answer: B
RNA-seq is a technology used for:
- A) Sequencing DNA
- B) Sequencing RNA
- C) Amplifying DNA
- D) Amplifying RNA
- Answer: B
Which method can be used to silence specific genes in a cell?
- A) Gene knockout
- B) RNA interference (RNAi)
- C) CRISPR-Cas9
- D) All of the above
- Answer: D
CRISPR technology can be used for:
- A) Gene editing
- B) Gene silencing
- C) Gene activation
- D) All of the above
- Answer: D
Which of the following is a tool for introducing mutations into RNA?
- A) Site-directed mutagenesis
- B) RNA editing
- C) RNA interference
- D) Reverse transcription
- Answer: B
RNA aptamers are:
- A) DNA molecules that bind specific targets
- B) RNA molecules that bind specific targets
- C) Proteins that bind specific RNA molecules
- D) Enzymes that degrade RNA
- Answer: B
Clinical and Experimental Applications of RNA
mRNA vaccines, such as those for COVID-19, work by:
- A) Delivering a live virus to stimulate an immune response
- B) Delivering mRNA that encodes a viral protein
- C) Delivering DNA that encodes a viral protein
- D) Delivering antibodies against the virus
- Answer: B
RNA therapeutics can be used to:
- A) Replace defective genes
- B) Silence disease-causing genes
- C) Enhance immune responses
- D) All of the above
- Answer: D
Which type of RNA is often used as a biomarker for disease?
- A) mRNA
- B) miRNA
- C) tRNA
- D) rRNA
- Answer: B
Antisense RNA therapy works by:
- A) Encoding for therapeutic proteins
- B) Complementing and binding to specific mRNA to block translation
- C) Enhancing mRNA stability
- D) Catalyzing RNA synthesis
- Answer: B
RNA molecules that can fold into complex three-dimensional structures are:
- A) Only found in prokaryotes
- B) Called ribozymes
- C) Only found in eukaryotes
- D) Incapable of catalytic activity
- Answer: B
Which of the following is a limitation of RNA-based therapeutics?
- A) Low specificity
- B) High stability in the body
- C) Potential for rapid degradation
- D) None of the above
- Answer: C
RNA-binding proteins play a critical role in:
- A) DNA replication
- B) RNA splicing
- C) Translation initiation
- D) All of the above
- Answer: D
Which RNA virus is known for causing the flu?
- A) HIV
- B) Influenza virus
- C) Hepatitis B virus
- D) Epstein-Barr virus
- Answer: B
RNA interference (RNAi) has been used to:
- A) Study gene function
- B) Develop therapeutics
- C) Engineer crops with desirable traits
- D) All of the above
- Answer: D
In situ hybridization (ISH) is used to:
- A) Measure protein levels
- B) Detect specific RNA sequences within tissues
- C) Sequence RNA
- D) Clone genes
- Answer: B
RNA viruses replicate by:
- A) Using the host's DNA polymerase
- B) Using their own RNA-dependent RNA polymerase
- C) Integrating into the host's genome
- D) Using the host's ribosomes to make RNA
- Answer: B
Which of the following is NOT an RNA virus?
- A) HIV
- B) Influenza virus
- C) Hepatitis C virus
- D) Hepatitis B virus
- Answer: D
Which enzyme transcribes HIV’s RNA genome into DNA?
- A) RNA polymerase
- B) DNA polymerase
- C) Reverse transcriptase
- D) Helicase
- Answer: C
RNA splicing occurs in:
- A) The cytoplasm
- B) The nucleus
- C) The ribosome
- D) The mitochondria
- Answer: B
The Central Dogma of molecular biology describes:
- A) DNA to RNA to protein
- B) RNA to DNA to protein
- C) Protein to RNA to DNA
- D) DNA to protein to RNA
- Answer: A
The coding sequence of a gene is typically found in:
- A) Exons
- B) Introns
- C) Promoters
- D) Enhancers
- Answer: A
The small interfering RNAs (siRNAs) are involved in:
- A) Protein synthesis
- B) Gene silencing
- C) DNA replication
- D) RNA splicing
- Answer: B
RNA molecules that have regulatory functions without coding for proteins are known as:
- A) rRNA
- B) tRNA
- C) ncRNA
- D) snRNA
- Answer: C
Which RNA modification enhances mRNA translation efficiency in eukaryotes?
- A) Splicing
- B) Addition of a 5' cap
- C) Polyadenylation
- D) Methylation
- Answer: B
Which of the following is an example of a ribonucleoprotein complex?
- A) DNA polymerase
- B) RNA polymerase
- C) Spliceosome
- D) Ribosome
- Answer: D



:max_bytes(150000):strip_icc():format(webp)/what-is-enzyme-structure-and-function-375555_FINAL-5a5a32ff18a24e13a6b985a7b64a34a4.png)



:max_bytes(150000):strip_icc()/electron-transport-chain-58e3be435f9b58ef7ed96112.jpg)