Protein Translation
1. Introduction
Protein translation is a complex and highly regulated process by which cells convert the genetic code in mRNA into a specific polypeptide chain. This process involves the precise coordination of various molecular components and stages, ensuring that proteins are synthesized accurately and efficiently.
2. Components of Translation
mRNA (Messenger RNA):
- Structure: Linear sequence with a 5' cap (7-methylguanylate cap) and a 3' poly(A) tail. The 5' cap protects the mRNA from degradation and aids in ribosome binding, while the poly(A) tail enhances stability and translation efficiency.
- Coding Region: Contains coding sequences organized into codons, each specifying a single amino acid.
tRNA (Transfer RNA):
- Structure: Cloverleaf secondary structure with an acceptor stem (where the amino acid attaches) and an anticodon loop (where the anticodon sequence binds to the mRNA codon).
- Aminoacylation: tRNAs are charged with amino acids by aminoacyl-tRNA synthetases. This process involves the enzyme recognizing both the tRNA and the specific amino acid.
Ribosome:
- Structure: Composed of two subunits. In prokaryotes, the small subunit is 30S and the large subunit is 50S, while in eukaryotes, the small subunit is 40S and the large subunit is 60S. The ribosome is a ribonucleoprotein complex that catalyzes peptide bond formation.
- Functional Sites:
- A (Aminoacyl) Site: Binds incoming aminoacyl-tRNA.
- P (Peptidyl) Site: Holds the tRNA with the growing polypeptide chain.
- E (Exit) Site: Where deacylated tRNA exits the ribosome.
Aminoacyl-tRNA Synthetases:
- Function: Catalyze the esterification of an amino acid to its corresponding tRNA. Each synthetase is specific to a single amino acid and its cognate tRNAs.
Translation Factors:
- Initiation Factors (IFs in prokaryotes, eIFs in eukaryotes): Facilitate ribosome assembly and mRNA binding.
- Elongation Factors (EFs in prokaryotes, eEFs in eukaryotes): Assist in the delivery of aminoacyl-tRNA to the ribosome and translocation of the ribosome along the mRNA.
- Release Factors (RFs in prokaryotes, eRFs in eukaryotes): Recognize stop codons and facilitate the release of the newly synthesized polypeptide.
3. Stages of Translation
3.1. Initiation
Formation of the Initiation Complex:
- Prokaryotes:
- The small ribosomal subunit (30S) binds to the mRNA at the Shine-Dalgarno sequence, which is complementary to a sequence in the 16S rRNA of the ribosome.
- The initiator tRNA carrying formylmethionine (fMet) binds to the start codon (AUG) in the P site.
- Initiation factors (IF1, IF2, and IF3) assist in this process.
- Eukaryotes:
- The small ribosomal subunit (40S) along with initiation factors (eIF1, eIF2, eIF3) binds to the 5' cap of the mRNA.
- The pre-initiation complex scans the mRNA for the AUG start codon.
- The initiator tRNA, charged with methionine, binds to the start codon, assisted by eIF5B.
- Prokaryotes:
Assembly of the Ribosome:
- The large ribosomal subunit (60S) joins the small subunit (40S), forming the functional 80S ribosome in eukaryotes, or the 70S ribosome in prokaryotes.
- The initiator tRNA occupies the P site, while the A site is primed for the next aminoacyl-tRNA.
3.2. Elongation
Codon Recognition:
- Aminoacyl-tRNA enters the A site of the ribosome. The correct tRNA is selected based on the codon-anticodon pairing. This process is mediated by elongation factor EF-Tu (prokaryotes) or eEF1A (eukaryotes), which ensures accuracy by binding GTP.
Peptide Bond Formation:
- The ribosome’s peptidyl transferase center, a ribozyme activity of the 23S rRNA in prokaryotes or the 28S rRNA in eukaryotes, catalyzes the formation of a peptide bond between the amino group of the amino acid in the A site and the carboxyl group of the amino acid in the P site.
Translocation:
- The ribosome moves along the mRNA by one codon. This movement is facilitated by elongation factor EF-G (prokaryotes) or eEF2 (eukaryotes) and involves the hydrolysis of GTP.
- The tRNA in the A site moves to the P site, and the tRNA in the P site moves to the E site before exiting the ribosome.
3.3. Termination
Stop Codon Recognition:
- When a stop codon (UAA, UAG, or UGA) reaches the A site, it is recognized by release factors (RF1 and RF2 in prokaryotes, eRF1 in eukaryotes).
- Release factors bind to the stop codon and promote the hydrolysis of the ester bond between the polypeptide chain and the tRNA in the P site.
Polypeptide Release:
- The newly synthesized polypeptide is released into the cytoplasm or the endoplasmic reticulum, depending on the cell type and protein destination.
Disassembly:
- The ribosomal subunits dissociate from the mRNA, and the tRNA molecules are released.
- The mRNA may be degraded or recycled for another round of translation.
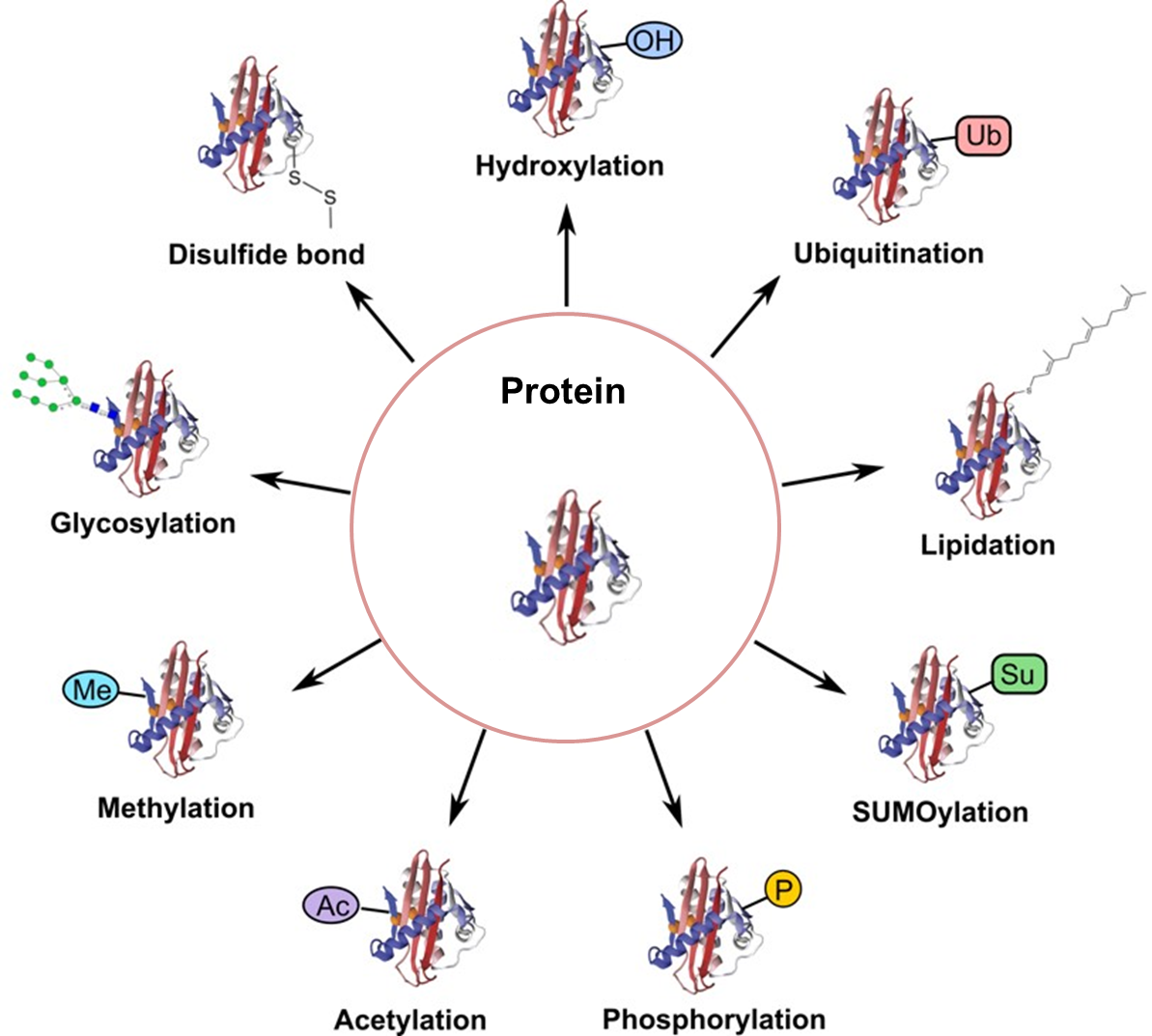
4. Post-Translational Modifications
Phosphorylation:
- Enzymatic addition of phosphate groups to serine, threonine, or tyrosine residues by kinases. This modification can alter enzyme activity, protein-protein interactions, and cellular localization.
Glycosylation:
- Addition of carbohydrate moieties to specific amino acid side chains, such as asparagine (N-linked glycosylation) or serine/threonine (O-linked glycosylation). Glycosylation affects protein folding, stability, and interactions.
Acetylation:
- Addition of acetyl groups to lysine residues by acetyltransferases. Acetylation can influence gene expression, protein stability, and protein-protein interactions.
Cleavage:
- Proteolytic cleavage by specific proteases can activate or inactivate proteins, remove signal sequences, or generate functional peptides from larger precursors.
5. Regulation of Translation
mRNA Stability:
- Decay Mechanisms: mRNA stability is regulated by factors such as the presence of AU-rich elements (AREs) in the 3' UTR, which are bound by decay-promoting proteins.
- Translational Repression: Binding of regulatory proteins or microRNAs (miRNAs) to the 3' UTR can inhibit translation or promote degradation.
Initiation Factors:
- Phosphorylation: Phosphorylation of initiation factors (e.g., eIF2α) can inhibit translation initiation in response to stress or nutrient deprivation.
- Regulatory Proteins: Specific proteins can modulate the activity of initiation factors and influence translation rates.
Global Regulation:
- Stress Response: Cellular stress responses (e.g., heat shock, oxidative stress) can alter translation through pathways like the integrated stress response (ISR), which involves eIF2α phosphorylation and inhibition of global translation.
- Nutrient Availability: Translation regulation in response to nutrient availability is mediated by signaling pathways such as the mTOR pathway, which affects the activity of translation initiation factors.
6. Translation in Prokaryotes vs. Eukaryotes
Prokaryotes:
- Coupled Transcription and Translation: In prokaryotes, translation begins while the mRNA is still being synthesized (coupled transcription and translation).
- Shine-Dalgarno Sequence: The small ribosomal subunit binds to the Shine-Dalgarno sequence in the mRNA, facilitating translation initiation.
Eukaryotes:
- Spatial Separation: Transcription occurs in the nucleus, and translation occurs in the cytoplasm. mRNA undergoes processing (5' capping, splicing, and polyadenylation) before translation.
- 5' Cap and Poly(A) Tail: The 5' cap and poly(A) tail enhance mRNA stability and translation efficiency. The cap-binding complex and poly(A)-binding proteins play key roles in initiation.
Please, give your suggetions in comment section.
Follow our blogs for more updates and protocols: https://iammolecularbiologist.blogspot.com/







0 Comments:
Post a Comment