Function of RNA
1. Introduction
RNA (ribonucleic acid) plays crucial roles in cellular processes, beyond merely acting as an intermediary between DNA and proteins. RNA functions are diverse and involve complex regulatory and catalytic activities. Understanding RNA's roles helps in comprehending how cells maintain homeostasis, respond to environmental signals, and regulate gene expression.
2. Types of RNA and Their Functions
2.1. Messenger RNA (mRNA)
Function:
- Genetic Information Transfer: Carries genetic information from DNA in the nucleus to ribosomes in the cytoplasm, where it directs protein synthesis.
- Translation Template: Specifies the sequence of amino acids in proteins through its codons.
Structure:
- 5' Cap: A 7-methylguanylate cap structure is added co-transcriptionally. It protects mRNA from exonucleolytic degradation, facilitates nuclear export, and is crucial for efficient translation initiation.
- 5' UTR: Contains regulatory elements that influence translation initiation and mRNA stability. Examples include upstream open reading frames (uORFs) that can modulate translation efficiency.
- Coding Region: Contains exons (coding sequences) that are translated into proteins. The sequence of codons determines the amino acid sequence of the polypeptide.
- 3' UTR: Contains regulatory sequences that control mRNA stability, localization, and translation efficiency. Includes elements like AU-rich elements (AREs) and binding sites for microRNAs (miRNAs).
Processing:
- Capping: Addition of the 5' cap is essential for mRNA stability, translation initiation, and splicing.
- Splicing: Involves the removal of introns and joining of exons. The spliceosome, a complex of snRNAs and proteins, facilitates this process.
- Polyadenylation: Addition of a poly(A) tail at the 3' end enhances mRNA stability and translation by protecting against exonucleolytic degradation and facilitating translation initiation.

2.2. Transfer RNA (tRNA)
Function:
- Amino Acid Transport: Transfers specific amino acids to the ribosome, where they are incorporated into the polypeptide chain.
- Codon-Anticodon Matching: Ensures the correct amino acid is added by matching its anticodon with the mRNA codon.
Structure:
- Anticodon Loop: A sequence of three nucleotides that base-pairs with a complementary mRNA codon.
- Acceptor Stem: The 3' end of the tRNA, where the amino acid is covalently attached by aminoacyl-tRNA synthetases.
Aminoacylation:
- Aminoacyl-tRNA Synthetases: These enzymes charge tRNA with amino acids using ATP. There are 20 synthetases, one for each amino acid, ensuring specificity through the recognition of tRNA and amino acid.
2.3. Ribosomal RNA (rRNA)
Function:
- Ribosome Structure: rRNA forms the core structure of ribosomes, providing a scaffold for ribosomal proteins and facilitating ribosome assembly.
- Catalytic Activity: The ribosome’s peptidyl transferase activity, which catalyzes peptide bond formation, is attributed to the 23S rRNA (prokaryotes) and 28S rRNA (eukaryotes).
Structure:
- Large Subunit rRNA: Includes 23S rRNA (prokaryotes) or 28S rRNA (eukaryotes), involved in peptide bond formation.
- Small Subunit rRNA: Includes 16S rRNA (prokaryotes) or 18S rRNA (eukaryotes), involved in mRNA binding and decoding.
2.4. Small Nuclear RNA (snRNA)
Function:
- Splicing: SnRNAs are key components of the spliceosome, which facilitates the removal of introns and the joining of exons in pre-mRNA.
- RNA Processing: Assist in various aspects of RNA processing, including the regulation of splicing and the export of mRNA from the nucleus.
Structure:
- U1, U2, U4, U5, U6 snRNAs: Each plays a specific role in splicing. For example, U1 snRNA recognizes the 5' splice site, while U2 snRNA binds to the branch point sequence.
2.5. MicroRNA (miRNA) and Small Interfering RNA (siRNA)
Function:
- Gene Silencing: miRNAs and siRNAs regulate gene expression post-transcriptionally by guiding the RNA-induced silencing complex (RISC) to target mRNAs, leading to their degradation or translational repression.
- RNA Interference (RNAi): A mechanism for gene silencing, where siRNAs guide the RISC to target and degrade complementary mRNA sequences.
Structure:
- miRNAs: Typically 21-23 nucleotides long, derived from hairpin structures in primary miRNA transcripts. Processed by Drosha and Dicer before being incorporated into RISC.
- siRNAs: Generally 20-25 nucleotides long, derived from long double-stranded RNA precursors. Processed by Dicer and incorporated into RISC.
2.6. Long Non-Coding RNA (lncRNA)
Function:
- Regulation of Gene Expression: lncRNAs can modulate gene expression at multiple levels, including transcription, splicing, and translation.
- Epigenetic Regulation: Interact with chromatin-modifying complexes and transcription factors, affecting chromatin structure and gene accessibility.
Structure:
- Varied Lengths: Typically longer than 200 nucleotides. Can be linear or have complex secondary structures.
- Diverse Functions: Include gene silencing, X-chromosome inactivation (e.g., XIST lncRNA), and regulation of developmental processes.
2.7. Small Nucleolar RNA (snoRNA)
Function:
- rRNA Modification: SnoRNAs guide the chemical modification of rRNA, such as methylation and pseudouridylation, which are critical for ribosome function and stability.
Structure:
- Box C/D and Box H/ACA Motifs: Conserved sequence motifs in snoRNAs that are essential for guiding specific rRNA modifications. Box C/D snoRNAs guide 2'-O-methylation, while Box H/ACA snoRNAs guide pseudouridylation.
2.8. Transfer-Messenger RNA (tmRNA)
Function:
- Rescue of Stalled Ribosomes: tmRNA rescues ribosomes stalled on defective mRNA by providing a template for the synthesis of a peptide tag that directs the incomplete protein for degradation.
Structure:
- Hybrid tRNA-mRNA Structure: Contains both tRNA-like and mRNA-like sequences. The tRNA-like part allows tmRNA to bind to the ribosome, while the mRNA-like part provides the sequence for tagging defective proteins.
3. RNA Synthesis and Processing
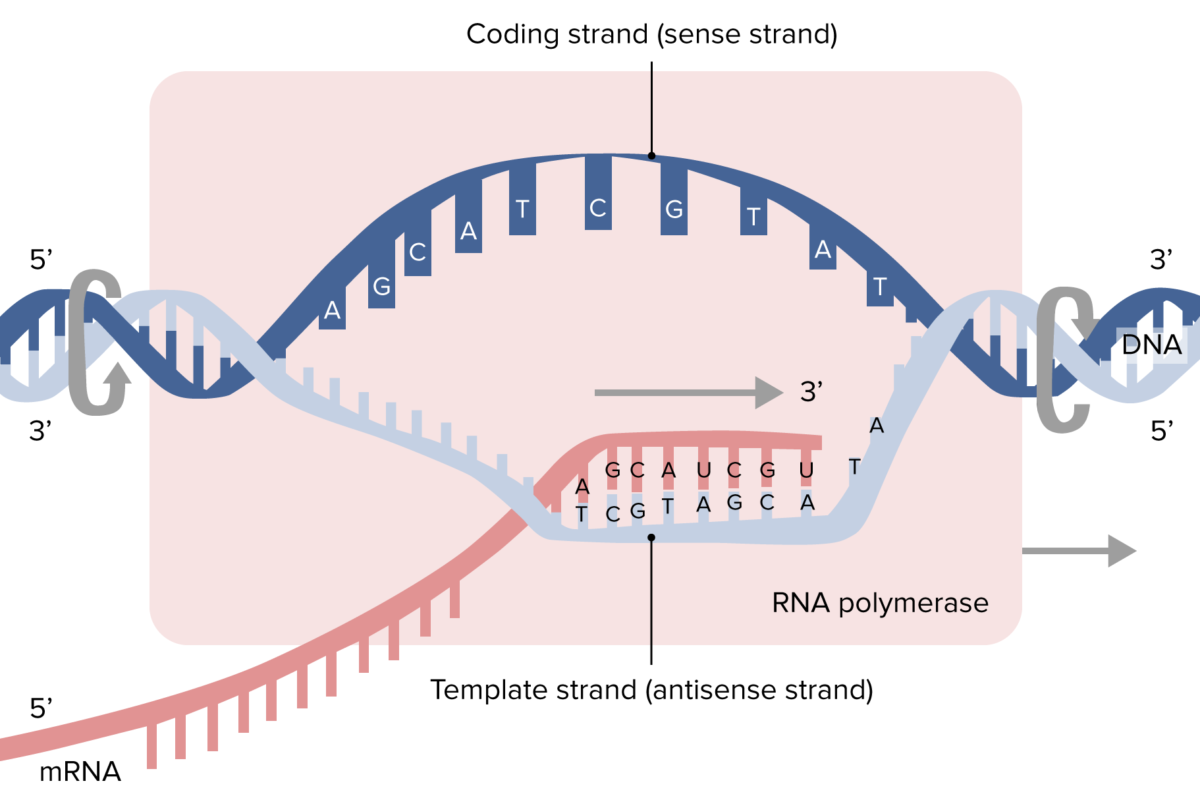
Transcription:
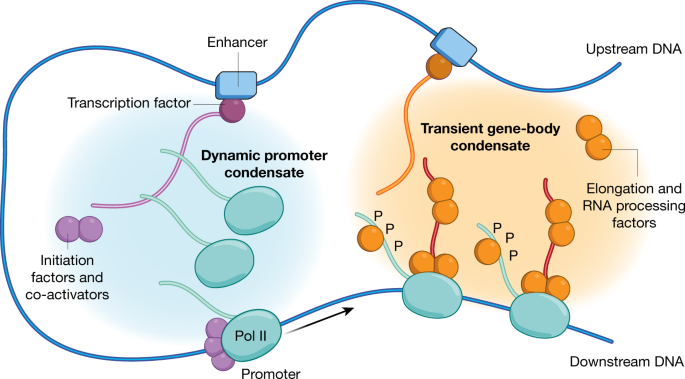
- Initiation: RNA polymerase binds to the promoter region of a gene, which is recognized by specific transcription factors. In eukaryotes, this involves general transcription factors and RNA polymerase II.
- Elongation: RNA polymerase synthesizes RNA in the 5' to 3' direction, unwinding the DNA and adding nucleotides complementary to the DNA template strand.
- Termination: In prokaryotes, termination occurs when RNA polymerase reaches a terminator sequence. In eukaryotes, termination involves cleavage of the pre-mRNA and the addition of a poly(A) tail.
Processing:
- Capping: Addition of the 5' cap involves the transfer of a 7-methylguanylate cap structure to the 5' end of the mRNA, which is crucial for mRNA stability and translation.
- Splicing: Involves the removal of introns and joining of exons through the spliceosome. Splicing is tightly regulated and can produce multiple protein isoforms through alternative splicing.
- Polyadenylation: Addition of a poly(A) tail, which involves cleavage of the pre-mRNA and subsequent addition of approximately 200 adenine residues. This modification protects the mRNA from degradation and aids in its export to the cytoplasm.
- Editing: RNA editing involves post-transcriptional modifications of RNA sequences, such as adenosine-to-inosine (A-to-I) editing, which can alter protein coding sequences and regulatory elements.
4. RNA Regulation
RNA Stability:
- Decapping and Degradation: The 5' cap can be removed by decapping enzymes, followed by exonucleolytic degradation of the mRNA. Degradation pathways include the exosome complex and nonsense-mediated decay (NMD).
- mRNA Surveillance: Mechanisms such as NMD ensure the elimination of faulty mRNA transcripts that contain premature stop codons.
RNA Interference (RNAi):
- Mechanism: Small RNA molecules (miRNAs and siRNAs) guide the RISC to target mRNAs based on complementary sequences. This results in mRNA degradation or inhibition of translation, modulating gene expression.
- Applications: RNAi is used in functional genomics to silence specific genes and investigate their functions. It has therapeutic potential for targeting disease-causing genes.
Alternative Splicing:
- Splice Variants: Pre-mRNA can be spliced in different ways to produce multiple protein isoforms from a single gene. Alternative splicing is regulated by splicing factors and can affect gene function and diversity.
5. RNA in Cellular Processes
Protein Synthesis:
- Translation: mRNA is translated into proteins by ribosomes with the help of tRNA and rRNA. The process involves initiation, elongation, and termination, and is tightly regulated to ensure accuracy.
- Ribosome Function: The ribosome's rRNA components provide the catalytic activity for peptide bond formation and facilitate the proper folding of the polypeptide chain.
Gene Regulation:
- Transcriptional Regulation: lncRNAs, small RNAs, and regulatory elements in mRNA (e.g., 5' UTRs) modulate gene expression by influencing transcriptional and post-transcriptional processes.
- Epigenetic Regulation: RNA molecules can interact with chromatin-modifying complexes to influence gene accessibility and expression, contributing to epigenetic regulation.
Cellular Responses:
- Stress Responses: RNA molecules, including small RNAs and lncRNAs, play roles in cellular responses to stress, such as heat shock or oxidative stress, by modulating gene expression and protein synthesis.
RNA Modifications:
- Chemical Modifications: RNA molecules undergo various chemical modifications, such as methylation and pseudouridylation, which affect their stability, structure, and function.
Please, write your suggetions in comment section.
Follow our blogs for more updates and protocols: https://iammolecularbiologist.blogspot.com/
Follow us on WhatsApp: https://chat.whatsapp.com/I30OdffWlJhGABXsTmKo95







0 Comments:
Post a Comment