Central Dogma of Molecular Biology
The Central Dogma of Molecular Biology explains the flow of genetic information within a biological system. It is the process by which the information in genes flows into proteins: DNA → RNA → Protein.
DNA Replication (for more details refer DNA (Deoxyribonucleic Acid) Chapter-5)
Mechanisms of Replication
- Origin of Replication: The specific sequence at which replication begins. In prokaryotes, there is usually a single origin of replication, while eukaryotes have multiple origins.
- Replication Fork: The Y-shaped structure formed during DNA replication where the DNA is split into two strands and replication occurs.
- Leading and Lagging Strand Synthesis: DNA replication is semi-discontinuous. The leading strand is synthesized continuously, while the lagging strand is synthesized in short segments called Okazaki fragments.
Key Enzymes and Proteins
- DNA Polymerases: Enzymes that synthesize new DNA strands by adding nucleotides to a pre-existing strand.
- Helicase: Unwinds the DNA double helix at the replication fork.
- Primase: Synthesizes RNA primers that provide a starting point for DNA polymerase.
- Ligase: Joins Okazaki fragments on the lagging strand by forming phosphodiester bonds.
- Single-Strand Binding Proteins (SSBs): Stabilize single-stranded DNA and prevent it from re-annealing or forming secondary structures.
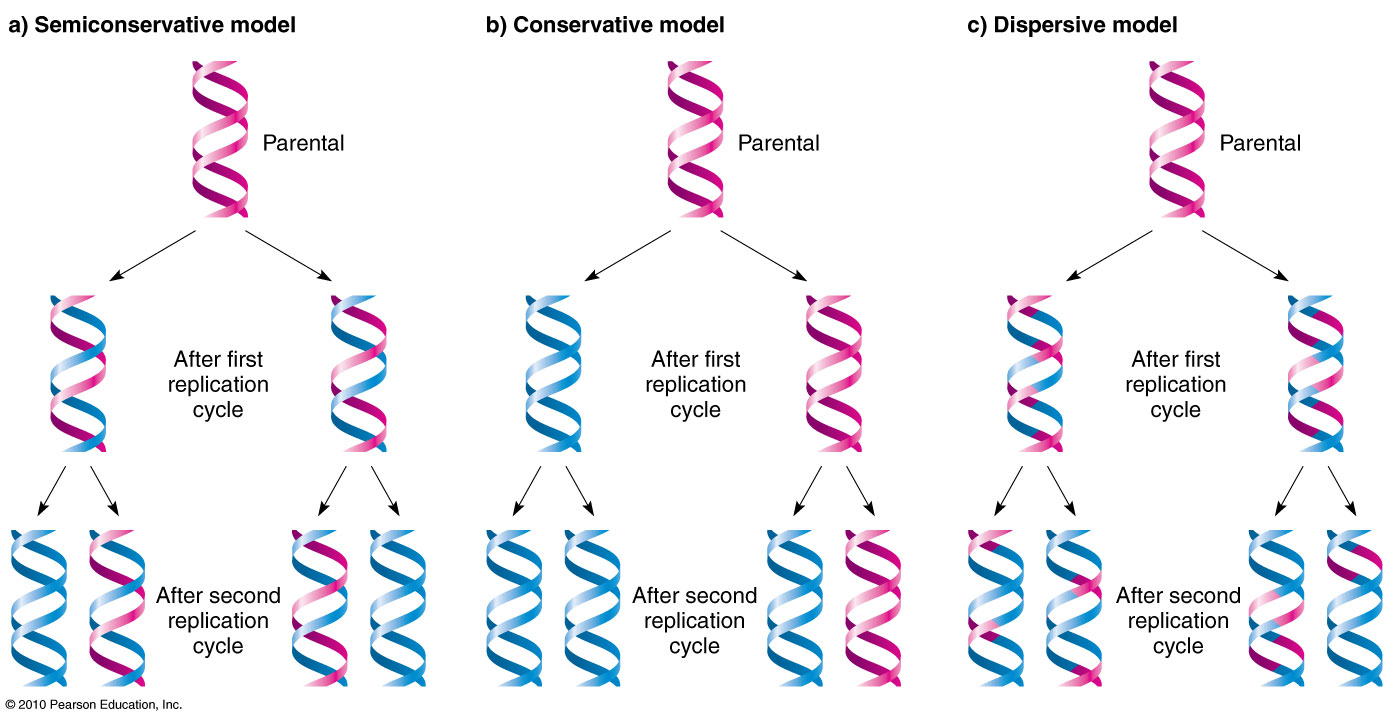
Replication Models
- Semi-Conservative Model: Each of the two daughter DNA molecules contains one old strand and one newly synthesized strand.
- Conservative Model: The original DNA molecule is conserved, and a completely new molecule is synthesized.
- Dispersive Model: Parental and newly synthesized DNA segments are interspersed in both strands following replication.
Proofreading and Repair (for more details refer DNA (Deoxyribonucleic Acid) Chapter-6)
- Proofreading Activity of DNA Polymerase: DNA polymerase has a 3' to 5' exonuclease activity that corrects errors during DNA synthesis.
- Mismatch Repair: Corrects errors missed by proofreading.
- Excision Repair: Removes and replaces damaged DNA, such as thymine dimers caused by UV light.
Transcription
RNA Synthesis
- Initiation: RNA polymerase binds to the promoter region of DNA and begins RNA synthesis.
- Elongation: RNA polymerase moves along the DNA template, synthesizing RNA in the 5' to 3' direction.
- Termination: RNA synthesis stops when RNA polymerase reaches a termination signal in the DNA.
Types of RNA
- mRNA (Messenger RNA): Carries the genetic code from DNA to the ribosome for protein synthesis.
- tRNA (Transfer RNA): Brings amino acids to the ribosome during translation.
- rRNA (Ribosomal RNA): Combines with proteins to form ribosomes.
- snRNA (Small Nuclear RNA): Involved in mRNA splicing.
- miRNA (Micro RNA): Regulates gene expression by interfering with mRNA.
Transcription in Prokaryotes vs Eukaryotes
- Prokaryotes: Transcription and translation occur simultaneously in the cytoplasm. Sigma factors help RNA polymerase recognize the promoter.
- Eukaryotes: Transcription occurs in the nucleus, and RNA undergoes extensive processing before translation. Transcription factors and RNA polymerase II are involved in initiation.
Post-Transcriptional Modifications
- 5' Capping: Addition of a modified guanine nucleotide to the 5' end of the mRNA.
- Polyadenylation: Addition of a poly(A) tail to the 3' end of the mRNA.
- Splicing: Removal of introns and joining of exons to produce mature mRNA.
- RNA Editing: Alteration of nucleotide sequences in RNA.
Translation
Mechanism of Translation
- Initiation: The small ribosomal subunit binds to the mRNA, and the initiator tRNA recognizes the start codon (AUG).
- Elongation: tRNAs bring amino acids to the ribosome, and peptide bonds form between amino acids.
- Termination: The ribosome reaches a stop codon, and the newly synthesized polypeptide is released.
Ribosome Structure and Function
- Ribosomal Subunits: Composed of a small and a large subunit that come together during translation.
- Sites within the Ribosome:
- A (Aminoacyl) Site: Holds the incoming tRNA with the next amino acid.
- P (Peptidyl) Site: Holds the tRNA with the growing polypeptide chain.
- E (Exit) Site: Where the empty tRNA exits the ribosome.
tRNA and Aminoacyl-tRNA Synthetase
- tRNA Structure: Cloverleaf structure with an anticodon at one end and an amino acid attachment site at the other.
- Charging of tRNA: Aminoacyl-tRNA synthetase attaches the correct amino acid to its corresponding tRNA.
Genetic Code
- Codon-Anticodon Pairing: The codon in mRNA pairs with the anticodon in tRNA.
- Start and Stop Codons: Start codon (AUG) initiates translation, while stop codons (UAA, UAG, UGA) terminate it.
- Wobble Hypothesis: The third base in a codon can pair with multiple bases in the anticodon, allowing for flexibility.
Post-Translational Modifications
- Phosphorylation: Addition of phosphate groups.
- Glycosylation: Addition of sugar moieties.
- Proteolytic Cleavage: Removal of specific peptide segments to activate the protein.
Gene Regulation (related to Central Dogma)
Operon Model in Prokaryotes
- Lac Operon: Regulates lactose metabolism. In the absence of lactose, the repressor binds to the operator, preventing transcription. In the presence of lactose, the repressor is inactivated, allowing transcription.
- Trp Operon: Regulates tryptophan synthesis. When tryptophan is abundant, it binds to the repressor, activating it to inhibit transcription.
Transcriptional Regulation in Eukaryotes
- Promoters, Enhancers, Silencers: DNA elements that regulate transcription.
- Transcription Factors: Proteins that bind to DNA and influence RNA polymerase binding and activity.
- Epigenetic Modifications:
- DNA Methylation: Addition of methyl groups to DNA, usually suppressing gene expression.
- Histone Modification: Addition or removal of chemical groups to histones, affecting chromatin structure and gene expression.
Key Experiments and Historical Context (for more details refer DNA (Deoxyribonucleic Acid) Chapter-3)
Griffith's Experiment (1928)
- Demonstrated the "transforming principle" by showing that non-virulent bacteria could become virulent when mixed with heat-killed virulent bacteria.
Avery-MacLeod-McCarty Experiment (1944)
- Identified DNA as the "transforming principle" by showing that DNA, not proteins or RNA, could transform bacteria.
Hershey-Chase Experiment (1952)
- Confirmed that DNA is the genetic material by using bacteriophages labeled with radioactive isotopes.
Meselson-Stahl Experiment (1958)
- Provided evidence for the semi-conservative model of DNA replication using isotopic labeling and density gradient centrifugation.
Please, write your suggestions in comment section.
Follow our blogs for more updates and protocols: https://iammolecularbiologist.blogspot.com/
Follow
us on WhatsApp: https://chat.whatsapp.com/I30OdffWlJhGABXsTmKo95






0 Comments:
Post a Comment